GUT MICROBIOTA DYSBIOSIS IN PEDIATRIC DIARRHEA: INSIGHTS FROM A PILOT STUDY IN DUHOK, KURDISTAN REGION OF IRAQ
Abstract
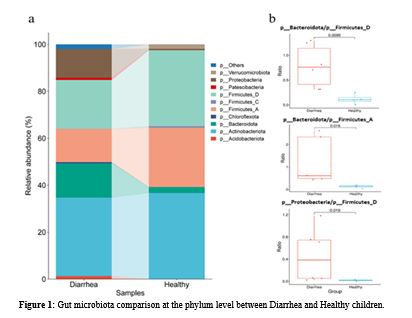
Diarrhea in children remains a significant global health burden and is closely linked to gut microbiota dysbiosis. However, data from Middle Eastern populations, particularly Iraq, remain limited. This study aims to characterize the change of gut microbiota associated with diarrhea in Iraqi children and to identify the potential of microbial imbalance by comparison with healthy controls. Stool samples were collected from 10 children (6 with diarrhea, 4 healthy controls) at Hevi Pediatric Teaching Hospital, Duhok, Iraq. Bacterial DNA was extracted and analyzed using 16S rRNA amplicon sequencing (V3–V4 region, Illumina NovaSeq). Sequence data were processed with QIIME2 software to assess taxonomic structure, α- and β-diversity. Comparisons between groups were performed using the Wilcoxon signed-rank test and the Metacoder Tree test. In this study, 241 bacterial taxa belonging to 11 phyla were identified. An enrichment in Proteobacteria and Bacteroidota was observed in cases of diarrhea, while the abundance of Firmicutes, alongside Ruminococcaceae, Lachnospiraceae, and Akkermansia muciniphila, was reduced. Notably, beneficial SCFA-producing genera, including Blautia, Faecalibacterium, and Bifidobacterium, were significantly reduced, while Enterobacteriaceae and Enterococcaceae were increased. Elevated ratios of Bacteroidota/Firmicutes and Proteobacteria/Firmicutes (p < 0.05) were also noted. Although no statistically significant differences were observed for α-diversity, β-diversity confirmed distinct clustering of both groups of diarrheal versus healthy microbiota. Diarrhea in Iraqi children was characterized by a shift of microbiome and a marked overgrowth of Proteobacteria, along with depletion of Firmicutes and loss of SCFA-producing genera, consistent with indicators of global microbial imbalance. However, the elevated Bacteroidota suggests region-specific indicators. These population-level variations may serve as biomarkers of imbalance. Further, larger, longitudinal, multi-omic studies are required to validate these findings.
Full text article
References
Ali, B. N. (2025). Molecular Detection Of Drug-Resistant Genes Among Clostridioides Difficile From Diarrheic Children in Duhok City-Iraq. Science Journal of University of Zakho, 13(2), 179–184.DOI: 10.25271/sjuoz.2025.13.2.1424
An, J., Kwon, H., & Kim, Y. J. (2023). The firmicutes/bacteroidetes ratio as a risk factor of breast cancer. Journal of Clinical Medicine, 12(6), 2216.https://doi.org/10.3390/jcm12062216
Biswas, L., Ibrahim, K. S., Li, X., Zhou, X., Zeng, Z., Craft, J., & Shu, X. (2021). Effect of a TSPO ligand on retinal pigment epithelial cholesterol homeostasis in high-fat fed mice, implication for age-related macular degeneration. Experimental Eye Research, 208(May), 108625. https://doi.org/10.1016/j.exer.2021.108625
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., & Holmes, S. P. (2016). DADA2: High-resolution sample inference from Illumina amplicon data. Nature Methods, 13(7), 581–583. https://doi.org/10.1038/nmeth.3869
Deering, K. E., Devine, A., O’Sullivan, T. A., Lo, J., Boyce, M. C., & Christophersen, C. T. (2019). Characterizing the composition of the pediatric gut microbiome: a systematic review. Nutrients, 12(1), 16.https://doi.org/10.3390/nu12010016
Du, Z., Li, J., Li, W., Fu, H., Ding, J., Ren, G., Zhou, L., Pi, X., & Ye, X. (2023). Effects of prebiotics on the gut microbiota in vitro associated with functional diarrhea in children. Frontiers in Microbiology, 14, 1233840. Doi: 10.3389/fmicb.2023.1233840.
Gallardo, P., Izquierdo, M., Vidal, R. M., Soto, F., Ossa, J. C., & Farfan, M. J. (2020). Gut microbiota-metabolome changes in children with diarrhea by diarrheagenic E. coli. Frontiers in Cellular and Infection Microbiology, 10, 485. Doi: 10.3389/fcimb.2020.00485.
George, S., Aguilera, X., Gallardo, P., Farfán, M., Lucero, Y., Torres, J. P., Vidal, R., & O’Ryan, M. (2022). Bacterial gut microbiota and infections during early childhood. Frontiers in Microbiology, 12, 793050. Doi: 10.3389/fmicb.2021.793050.
Hama-Ali, M. A., & Hasan, A. H. (2023). Profiling of Bacterial Species from Covid-19 Faecal Samples in Kurdistan Region-Iraq. Science Journal of University of Zakho, 11(1), 11–15. https://doi.org/10.25271/sjuoz.2022.10.4.1031.
Hasan, H. K., Yassin, N. A., & Eassa, S. H. (2020). Bacteriological and molecular characterization of diarrheagenic Escherichia coli pathotypes from children in Duhok City, Iraq. Science Journal of University of Zakho, 8(2), 52–57.https://doi.org/10.25271/sjuoz.2020.8.2.702.
Hernández, M., De Frutos, M., Rodríguez-Lázaro, D., López-Urrutia, L., Quijada, N. M., & Eiros, J. M. (2019). Fecal microbiota of toxigenic Clostridioides difficile-associated diarrhea. Frontiers in Microbiology, 9, 3331. Doi: 10.3389/fmicb.2018.03331.
Holmberg, S. M., Feeney, R. H., Prasoodanan PK, V., Puértolas-Balint, F., Singh, D. K., Wongkuna, S., Zandbergen, L., Hauner, H., Brandl, B., & Nieminen, A. I. (2024). The gut commensal Blautia mai.ntains colonic mucus function under low-fiber consumption through secretion of short-chain fatty acids. Nature Communications, 15(1), 3502. https://doi.org/10.1038/s41467-024-47594-w.
Iancu, M. A., Profir, M., Roşu, O. A., Ionescu, R. F., Cretoiu, S. M., & Gaspar, B. S. (2023). Revisiting the intestinal microbiome and its role in diarrhea and constipation. Microorganisms, 11(9), 2177. https://doi.org/10.3390/microorganisms11092177.
Ibrahim, K. S., Bourwis, N., Dolan, S., Lang, S., Spencer, J., & Craft, J. A. (2021). Characterisation of gut microbiota of obesity and type 2 diabetes in a rodent model. Bioscience of Microbiota, Food and Health, 40(1), 65–74. https://doi.org/10.12938/bmfh.2019-031.
Ibrahim, K. S., Craft, J. A., Biswas, L., Spencer, J., & Shu, X. (2020). Etifoxine reverses weight gain and alters the colonic bacterial community in a mouse model of obesity. Biochemical Pharmacology, 180. https://doi.org/10.1016/j.bcp.2020.114151
Jin, D., Chen, C., Li, L., Lu, S., Li, Z., Zhou, Z., Jing, H., Xu, Y., Du, P., & Wang, H. (2013). Dynamics of fecal microbial communities in children with diarrhea of unknown etiology and genomic analysis of associated Streptococcus lutetiensis. BMC Microbiology, 13(1), 141. https://doi.org/10.1186/1471-2180-13-141.
Kadhim, F. J., Aziz, Z. S., & Ibrahim, K. S. (2025). Gut Microbiome Profiles in Colorectal Cancer Patients in Iraq. Microbiology Research, 16(1), 22. https://doi.org/10.3390/microbiolres16010022.
Kang, Z., Lu, M., Jiang, M., Zhou, D., & Huang, H. (2019). Proteobacteria acts as a pathogenic risk-factor for chronic abdominal pain and diarrhea in post-cholecystectomy syndrome patients: a gut microbiome metabolomics study. Medical Science Monitor: International Medical Journal of Experimental and Clinical Research, 25, 7312. https://doi.org/10.12659/MSM.915984.
Kieser, S., Sarker, S. A., Sakwinska, O., Foata, F., Sultana, S., Khan, Z., Islam, S., Porta, N., Combremont, S., & Betrisey, B. (2018). Bangladeshi children with acute diarrhoea show faecal microbiomes with increased Streptococcus abundance, irrespective of diarrhoea aetiology. Environmental Microbiology, 20(6), 2256–2269. https://doi.org/10.1111/1462-2920.14274.
Ku, J.-Y., Lee, M.-J., Jung, Y., Choi, H.-J., & Park, J. (2025). Changes in the gut microbiome due to diarrhea in neonatal Korean indigenous calves. Frontiers in Microbiology, 16, 1511430. Doi: 10.3389/fmicb.2025.1511430.
Li, Y., Xia, S., Jiang, X., Feng, C., Gong, S., Ma, J., Fang, Z., Yin, J., & Yin, Y. (2021). Gut microbiota and diarrhea: an updated review. Frontiers in Cellular and Infection Microbiology, 11, 625210. Doi: 10.3389/fcimb.2021.625210.
Schlechte, J., Zucoloto, A. Z., Yu, I., Doig, C. J., Dunbar, M. J., McCoy, K. D., & McDonald, B. (2023). Dysbiosis of a microbiota–immune metasystem in critical illness is associated with nosocomial infections. Nature Medicine, 29(4), 1017–1027. https://doi.org/10.1038/s41591-023-02243-5.
Stojanov, S., Berlec, A., & Štrukelj, B. (2020). The influence of probiotics on the firmicutes/bacteroidetes ratio in the treatment of obesity and inflammatory bowel disease. Microorganisms, 8(11), 1715. https://doi.org/10.3390/microorganisms8111715.
Sun, J., Du, L., Li, X., Zhong, H., Ding, Y., Liu, Z., & Ge, L. (2019). Identification of the core bacteria in rectums of diarrheic and non-diarrheic piglets. Scientific Reports, 9(1), 18675. https://doi.org/10.1038/s41598-019-55328-y.
Tesfaw, G., Siraj, D. S., Abdissa, A., Jakobsen, R. R., Johansen, Ø. H., Zangenberg, M., Hanevik, K., Mekonnen, Z., Langeland, N., & Bjørang, O. (2024). Gut microbiota patterns associated with duration of diarrhea in children under five years of age in Ethiopia. Nature Communications, 15(1), 7532. https://doi.org/10.1038/s41467-024-51464-w.
The, H. C., & Le, S.-N. H. (2022). Dynamic of the human gut microbiome under infectious diarrhea. Current Opinion in Microbiology, 66, 79–85.https://doi.org/10.1016/j.mib.2022.01.006.
Wu, Z.-L., Wei, R., Tan, X., Yang, D., Liu, D., Zhang, J., & Wang, W. (2022). Characterization of gut microbiota dysbiosis of diarrheic adult yaks through 16S rRNA gene sequences. Frontiers in Veterinary Science, 9, 946906. Doi: 10.3389/fvets.2022.946906.
Wu, Z., Xu, Q., Gu, S., Chen, Y., Lv, L., Zheng, B., Wang, Q., Wang, K., Wang, S., & Xia, J. (2022). Akkermansia muciniphila ameliorates Clostridioides difficile infection in mice by modulating the intestinal microbiome and metabolites. Frontiers in Microbiology, 13, 841920. https://doi.org/10.3389/fmicb.2022.841920.
Xi, L., Song, Y., Qin, X., Han, J., & Chang, Y.-F. (2021). Microbiome analysis reveals the dynamic alternations in gut microbiota of diarrheal giraffa camelopardalis. Frontiers in Veterinary Science, 8, 649372. Doi: 10.3389/fvets.2021.649372.
Zhuang, X., Tian, Z., Li, L., Zeng, Z., Chen, M., & Xiong, L. (2018). Fecal microbiota alterations associated with diarrhea-predominant irritable bowel syndrome. Frontiers in Microbiology, 9, 1600. Doi: 10.3389/fmicb.2018.01600.
Parks, D. H., Chaumeil, P. A., Mussig, A. J., Rinke, C., Chuvochina, M. and Hugenholtz, P., (2026). GTDB release 10: a complete and systematic taxonomy for 715 230 bacterial and 17 245 archaeal genomes. Nucleic Acids Research, 54(D1), pp.D743-D754. https://doi.org/10.1093/nar/gkaf1458.
McDonald, D., Jiang, Y., Balaban, M., Cantrell, K., Zhu, Q., Gonzalez, A., Morton, J.T., Nicolaou, G., Parks, D.H., Karst, S.M. and Albertsen, M., 2024. Greengenes2 unifies microbial data in a single reference tree. Nature biotechnology, 42(5), pp.715-718. https://doi.org/10.1038/s41587-023-01845-1.
Foster ZS, Sharpton TJ, Grünwald NJ. Metacoder: An R package for visualization and manipulation of community taxonomic diversity data. PLoS computational biology. 2017 Feb 21;13(2):e1005404. https://doi.org/10.1371/journal.pcbi.1005404.
Authors
Copyright (c) 2026 Ali Y. Saeed, Bakhtyar N. Ali, Delveen R. Ibrahim, Khalid S. Ibrahim, Oaikhena E. Enimie, Imran Khan, Amir Abdulmawjood, and Zahra Naeef Ayoub

This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
Authors who publish with this journal agree to the following terms:
- Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution License [CC BY-NC-SA 4.0] that allows others to share the work with an acknowledgment of the work's authorship and initial publication in this journal.
- Authors are able to enter into separate, additional contractual arrangements for the non-exclusive distribution of the journal's published version of the work, with an acknowledgment of its initial publication in this journal.
- Authors are permitted and encouraged to post their work online.